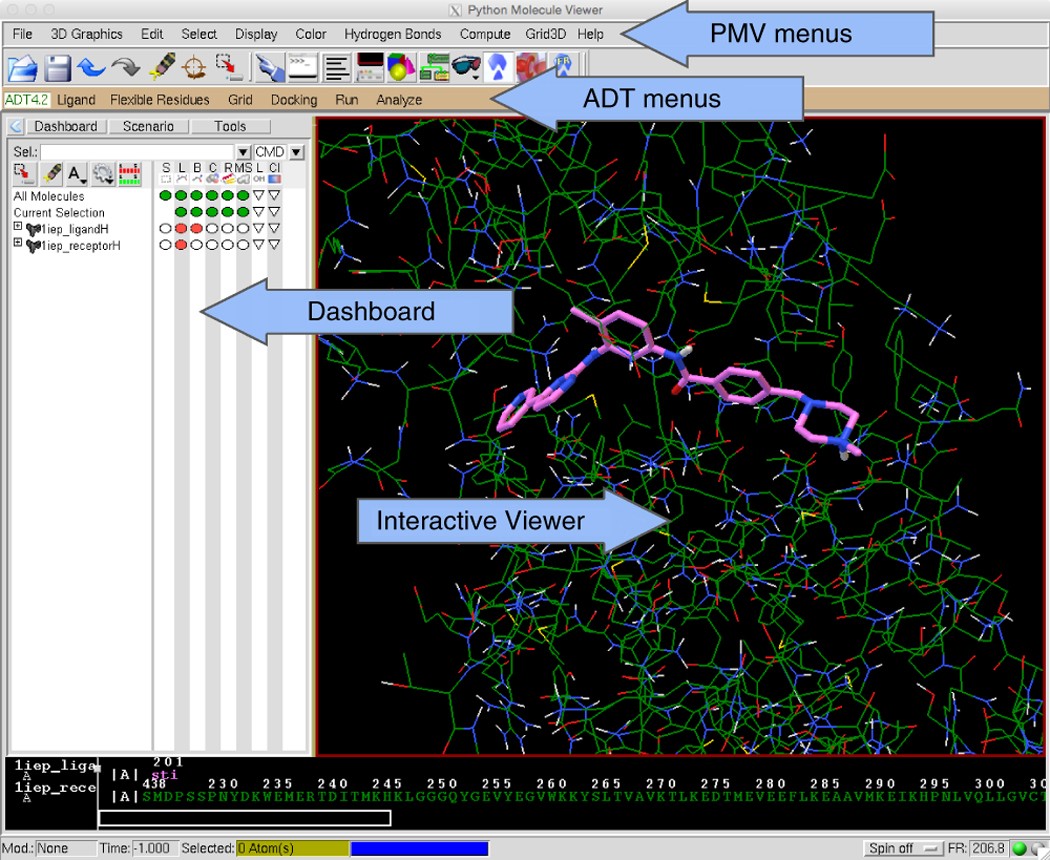
Computational protein–ligand docking and virtual drug screening with the AutoDock suite | Nature Protocols

Deep Docking: A Deep Learning Platform for Augmentation of Structure Based Drug Discovery | ACS Central Science

Benchmarking the Ability of Common Docking Programs to Correctly Reproduce and Score Binding Modes in SARS-CoV-2 Protease Mpro | Journal of Chemical Information and Modeling